pFC9 gel purification and pFC PCRs #
Recieved both SacI and primers in the mail this morning so today will be working on extracting large fragment of pFC9 after digestion with SacI and EcoRI for use in T7 initiation series cloning and PCR of various pFC8 and 9 plasmids for sequencing later this week.
pFC9 EcoRI and SacI digest #
Reagents and volumes are outlined in the spreadsheet at this link.
Expected results for both digests is shown below.
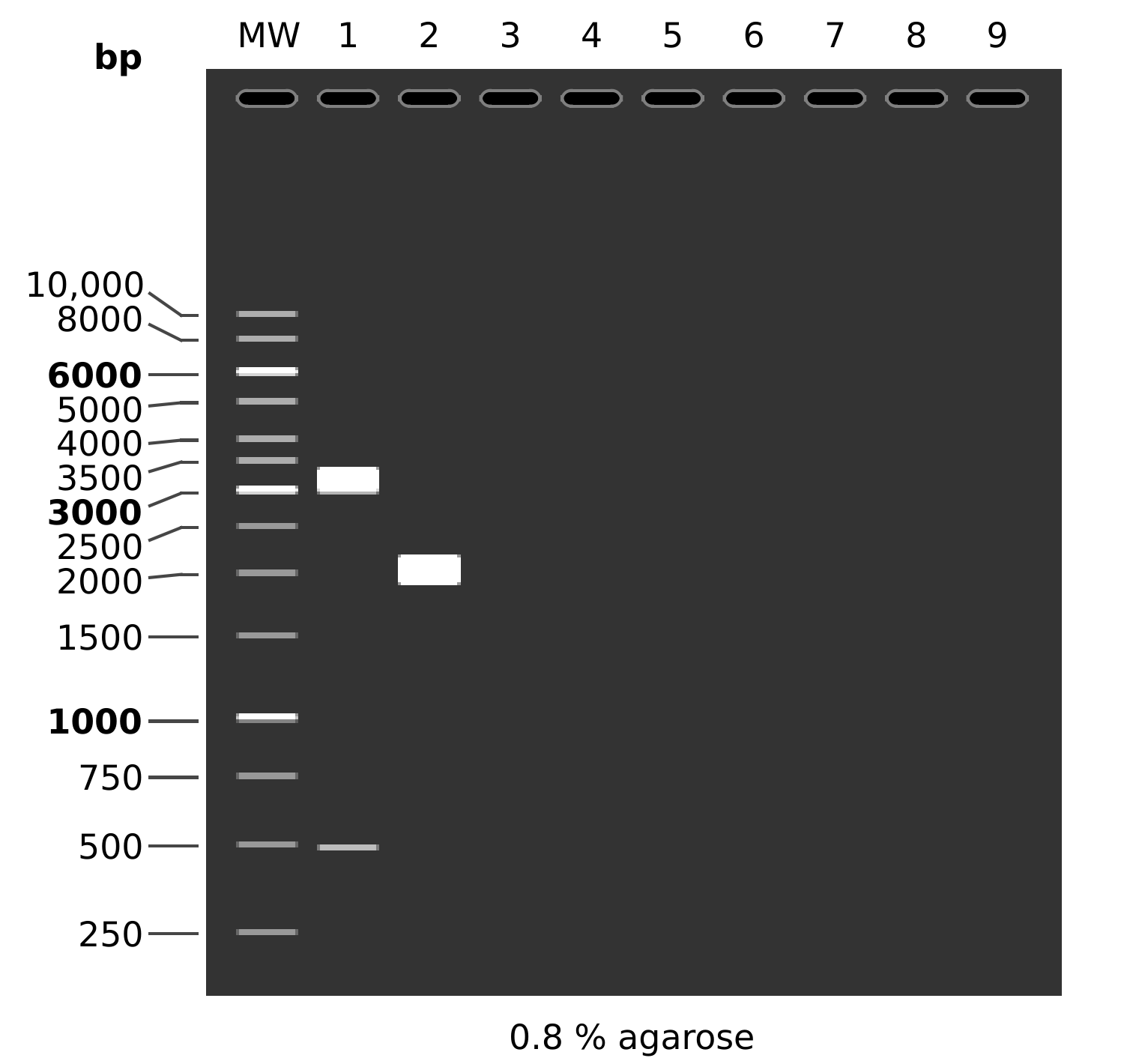
MW: GeneRuler™ 1 kb DNA Ladder
1: pFC9
SacI + EcoRI
1. 3099 bp
2. 490 bp
2: pFC9
1. 3589 bp
I would then excise the large fragment and purify using the Zymogen DNA agarose gel extraction kit reagents and protocol.
Digest 1 #
For first digest I used the buffer included with SacI which I thought was CutSmart but was actually rCurSmart, not sure if that is actually different in anyway or not.
Results #
Ran 0.08 agarose gel for one hour at 90 volts in TAE buffer.
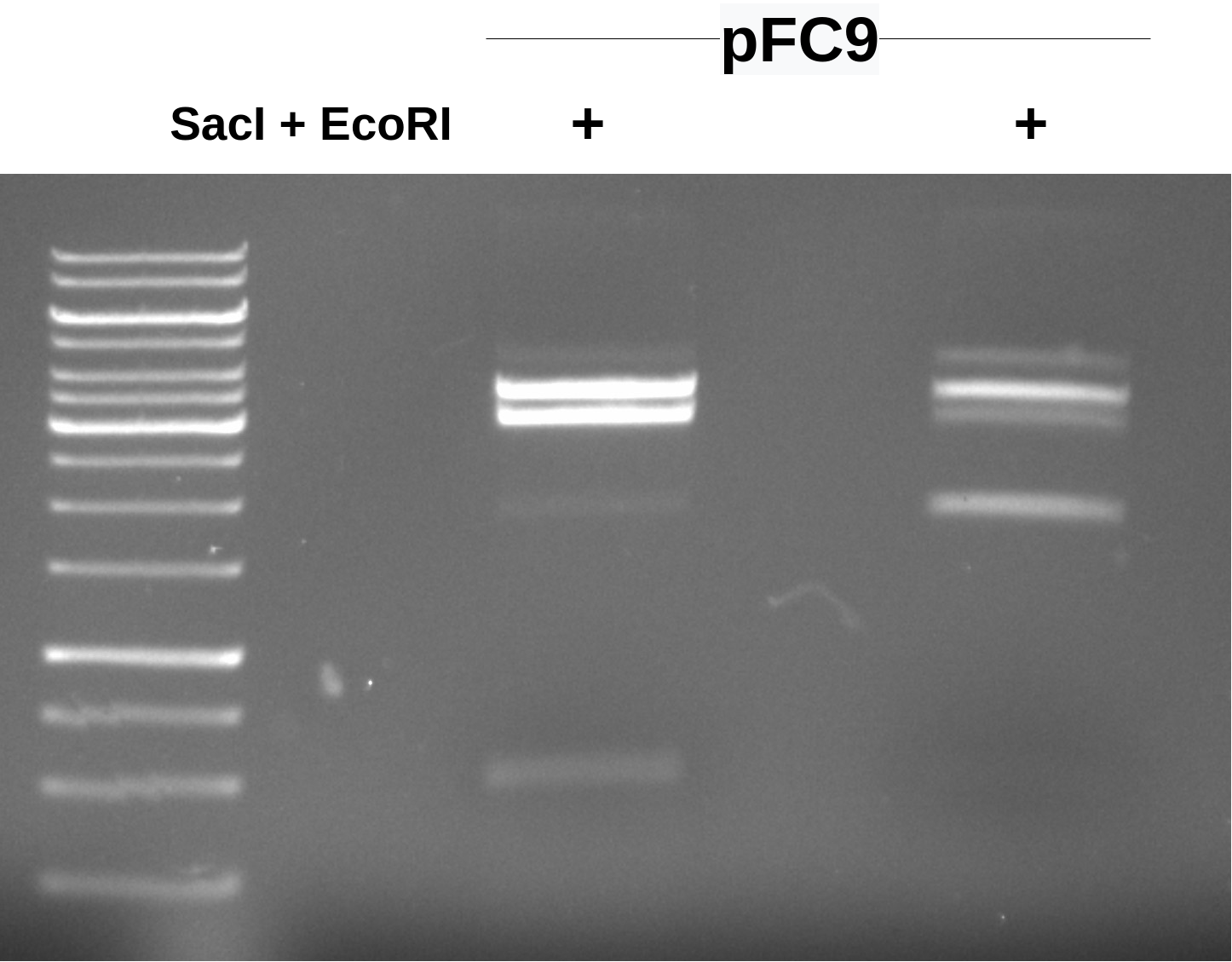
This mostly looks like pFC9 in so much that in lane 1 there are the two bands we would expect at ~3000 bp and at ~500 bp, although this 500 bp fragment is not present in the second digest (one enyme did not make it in?) and there is significant signal from a band of similar length around 3500-4000bp.
I am going to run this gel again and for longer to try and get better separation of the bands to make cutting out the pFC9 large fragment easier.
Digest 2 #
After band cutting I removed the band closet to the 3000 bp ladder marker.
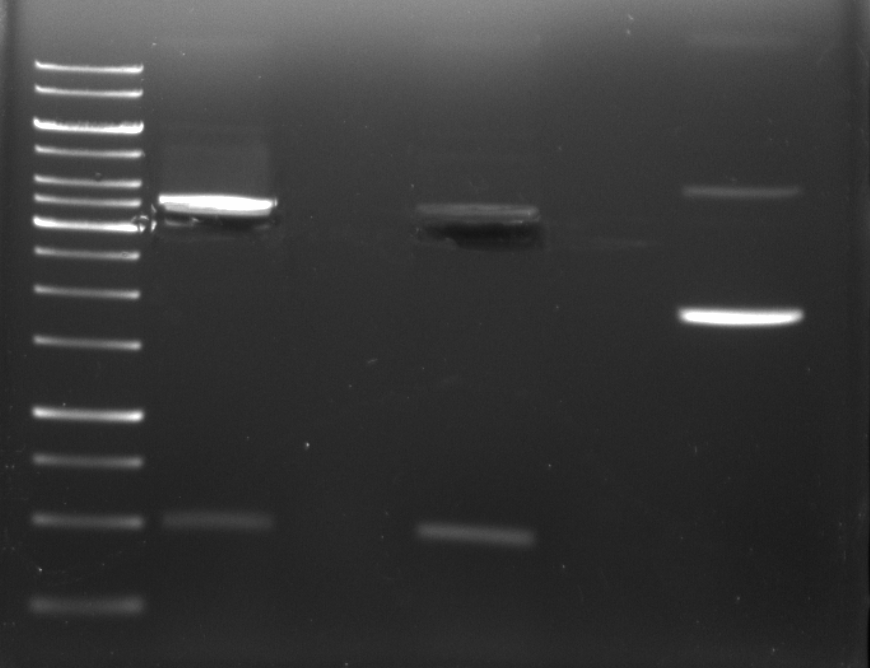
PCR #
PCR amplification in order to sequence pFC plasmids. More details on background and motivation for this assay are in notes from 7/29/21 and 7/28/21. All reagents, primers and concentrations are described in the spreadsheet at this link.
Results #
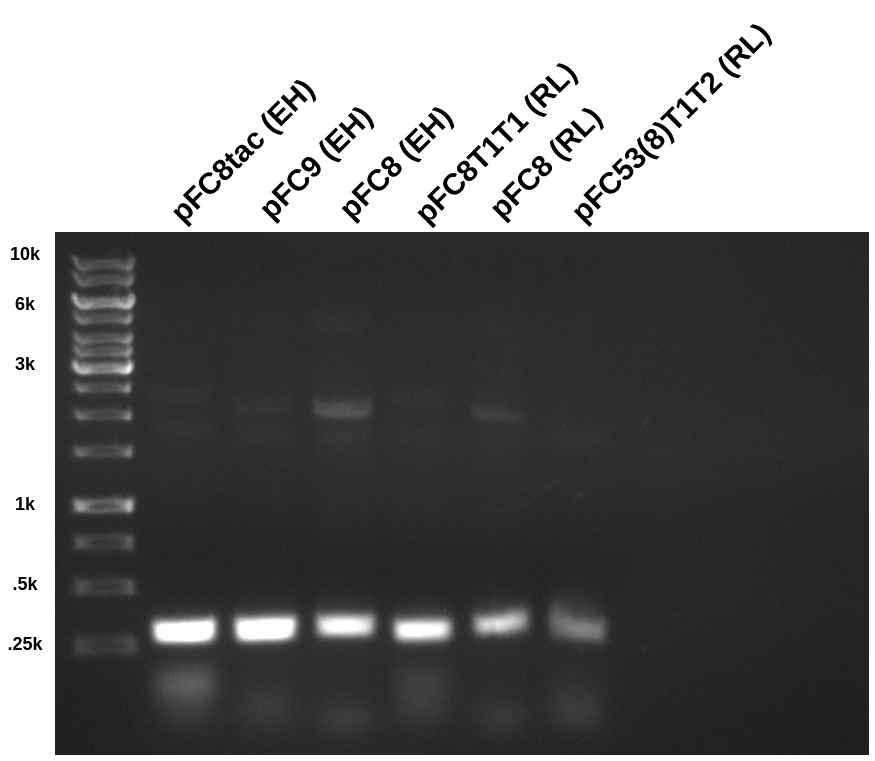
Followed up with agarose gel DNA extraction
| Sample name | Sample number | Assay |
|---|---|---|
| pFC8tac(EH) | 1 | PCR |
| pFC9(EH) | 2 | PCR |
| pFC8(EH) | 3 | PCR |
| pFC8T1T2(RL) | 4 | PCR |
| pFC8(RL) | 5 | PCR |
| pFC53T1T2(RL) | 6 | PCR |
| pFC9 | 7 | SacI EcoRI digest |
| pFC9 | 8 | SacI EcoRI digest |
Nanodrop result
| Sample | DNA concentration (ng/ul) | 260/280 | 260/230 |
|---|---|---|---|
| pFC8tac(EH) | -0.8 | ||
| pFC9(EH) | 12.7 | 1.43 | 0.321 |
| pFC8(EH) | 6.3 | 1.312 | 0.073 |
| pFC8T1T2(RL) | 4.1 | 1.141 | 0.117 |
| pFC8(RL) | -10.5 | 2.563 | -0.261 |
| pFC53T1T2(RL) | |||
| pFC9 | 2.2 | 1.0 | 0.106 |
| pFC9 | 8.5 | 1.689 | 0.179 |